Structure of the LCMV nucleoprotein provides a template for understanding arenavirus replication and immunosuppression.
West, B.R., Hastie, K.M., Saphire, E.O.(2014) Acta Crystallogr D Biol Crystallogr 70: 1764-1769
- PubMed: 24914986
- DOI: https://doi.org/10.1107/S1399004714007883
- Primary Citation of Related Structures:
4O6H, 4O6I - PubMed Abstract:
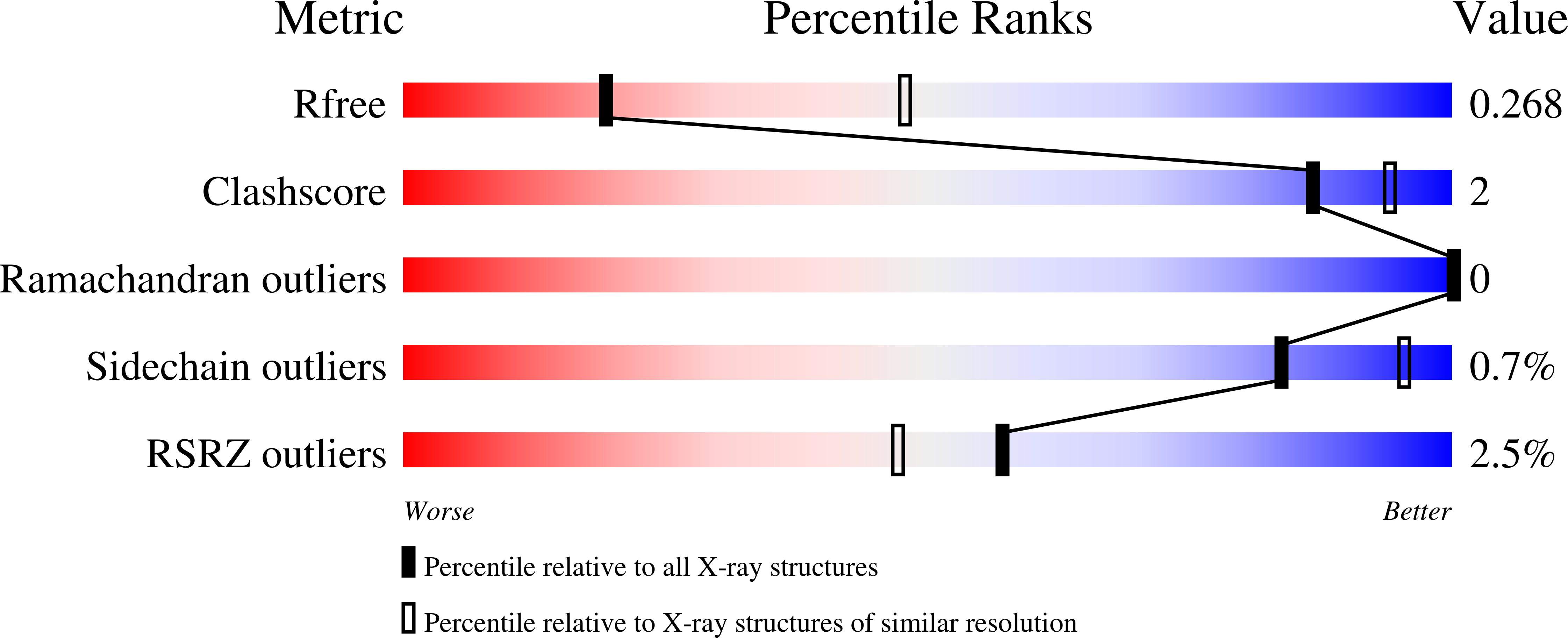
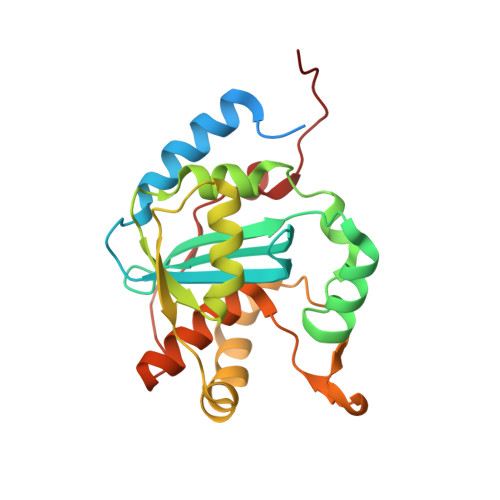
The X-ray crystal structure of the Lymphocytic choriomeningitis virus nucleoprotein C-terminal immunosuppressive domain (LCMV NPΔ340) was determined to 2.0 Å resolution. The structure indicates that LCMV NPΔ340, like the other structurally characterized arenaviral nucleoproteins, adopts the fold of an exonuclease. This structure provides a crucial three-dimensional template for functional exploration of the replication and immunosuppression of this prototypic arenavirus.
Organizational Affiliation:
Department of Immunology and Microbial Science, The Scripps Research Institute, La Jolla, CA 92037, USA.