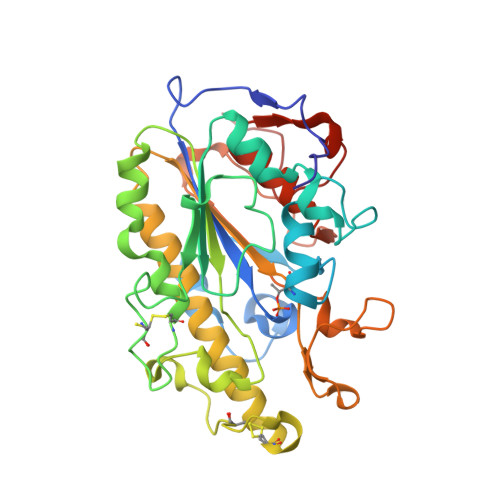
Substrate analog interaction with MCR-1 offers insight into the rising threat of the plasmid-mediated transferable colistin resistance.
Wei, P., Song, G., Shi, M., Zhou, Y., Liu, Y., Lei, J., Chen, P., Yin, L.(2018) FASEB J 32: 1085-1098
- PubMed: 29079699
- DOI: https://doi.org/10.1096/fj.201700705R
- Primary Citation of Related Structures:
5YLC, 5YLE, 5YLF - PubMed Abstract:
Colistin is considered a last-resort antibiotic against most gram-negative bacteria. Recent discoveries of a plasmid-mediated, transferable mobilized colistin-resistance gene ( mcr-1) on all continents have heralded the imminent emergence of pan-drug-resistant superbacteria. The inner-membrane protein MCR-1 can catalyze the transfer of phosphoethanolamine (PEA) to lipid A, resulting in colistin resistance. However, little is known about the mechanism, and few drugs exist to address this issue. We present crystal structures revealing the MCR-1 catalytic domain (cMCR-1) as a monozinc metalloprotein with ethanolamine (ETA) and d-glucose, respectively, thus highlighting 2 possible substrate-binding pockets in the MCR-1-catalyzed PEA transfer reaction. Mutation of the residues involved in ETA and d-glucose binding impairs colistin resistance in recombinant Escherichia coli containing full-length MCR-1. Partial analogs of the substrate are used for cocrystallization with cMCR-1, providing valuable information about the family of PEA transferases. One of the analogs, ETA, causes clear inhibition of polymyxin B resistance, highlighting its potential for drug development. These data demonstrate the crucial role of the PEA- and lipid A-binding pockets and provide novel insights into the structure-based mechanisms, important drug-target hot spots, and a drug template for further drug development to combat the urgent, rising threat of MCR-1-mediated antibiotic resistance.-Wei, P., Song, G., Shi, M., Zhou, Y., Liu, Y., Lei, J., Chen, P., Yin, L. Substrate analog interaction with MCR-1 offers insight into the rising threat of the plasmid-mediated transferable colistin resistance.
Organizational Affiliation:
State Key Laboratory of Virology, Hubei Key Laboratory of Cell Homeostasis, Department of Biochemistry and Molecular Biology, College of Life Sciences, Wuhan University, Wuhan, China.