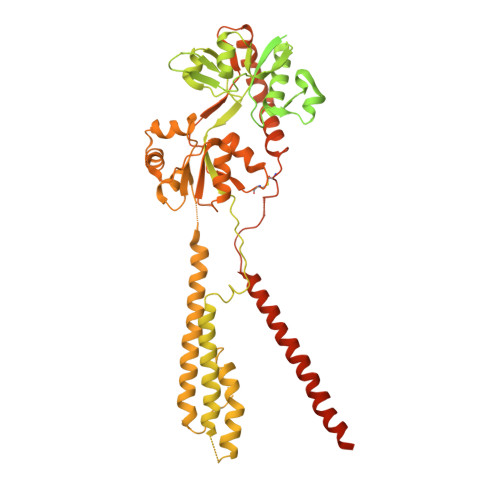
Architecture of fully occupied GluA2 AMPA receptor-TARP complex elucidated by cryo-EM.
Zhao, Y., Chen, S., Yoshioka, C., Baconguis, I., Gouaux, E.(2016) Nature 536: 108-111
- PubMed: 27368053
- DOI: https://doi.org/10.1038/nature18961
- Primary Citation of Related Structures:
5KK2 - PubMed Abstract:
Fast excitatory neurotransmission in the mammalian central nervous system is largely carried out by AMPA-sensitive ionotropic glutamate receptors. Localized within the postsynaptic density of glutamatergic spines, AMPA receptors are composed of heterotetrameric receptor assemblies associated with auxiliary subunits, the most common of which are transmembrane AMPA receptor regulatory proteins (TARPs). The association of TARPs with AMPA receptors modulates receptor trafficking and the kinetics of receptor gating and pharmacology. Here we report the cryo-electron microscopy (cryo-EM) structure of the homomeric rat GluA2 AMPA receptor saturated with TARP γ2 subunits, which shows how the TARPs are arranged with four-fold symmetry around the ion channel domain and make extensive interactions with the M1, M2 and M4 transmembrane helices. Poised like partially opened ‘hands’ underneath the two-fold symmetric ligand-binding domain (LBD) 'clamshells', one pair of TARPs is juxtaposed near the LBD dimer interface, whereas the other pair is near the LBD dimer-dimer interface. The extracellular ‘domains’ of TARP are positioned to not only modulate LBD clamshell closure, but also affect conformational rearrangements of the LBD layer associated with receptor activation and desensitization, while the TARP transmembrane domains buttress the ion channel pore.