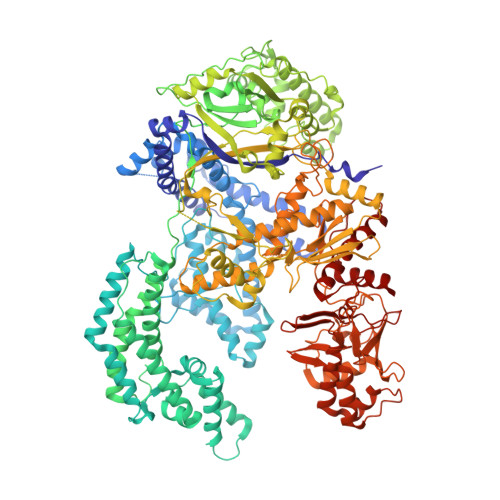
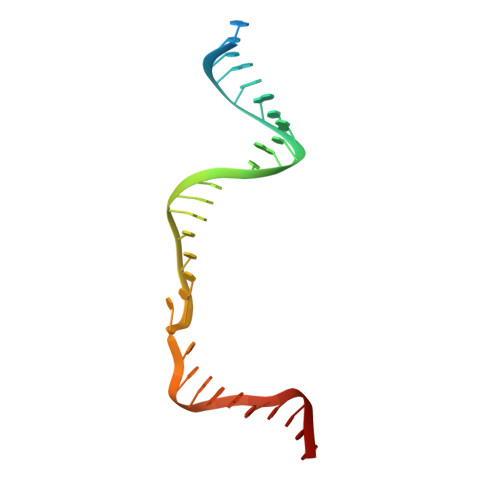
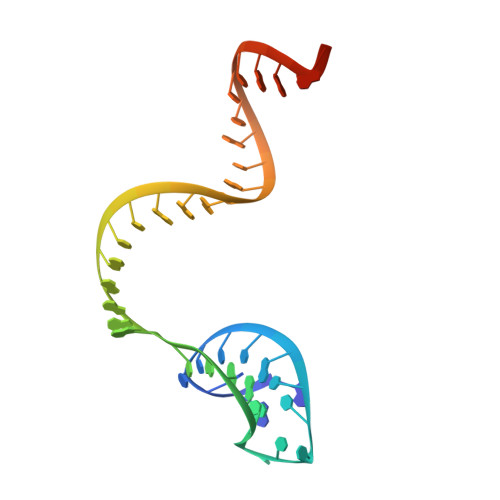Crystal Structure of Cpf1 in Complex with Guide RNA and Target DNA
Yamano, T., Nishimasu, H., Zetsche, B., Hirano, H., Slaymaker, I.M., Li, Y., Fedorova, I., Nakane, T., Makarova, K.S., Koonin, E.V., Ishitani, R., Zhang, F., Nureki, O.(2016) Cell 165: 949-962
- PubMed: 27114038
- DOI: https://doi.org/10.1016/j.cell.2016.04.003
- Primary Citation of Related Structures:
5B43 - PubMed Abstract:
Cpf1 is an RNA-guided endonuclease of a type V CRISPR-Cas system that has been recently harnessed for genome editing. Here, we report the crystal structure of Acidaminococcus sp. Cpf1 (AsCpf1) in complex with the guide RNA and its target DNA at 2.8 Å resolution. AsCpf1 adopts a bilobed architecture, with the RNA-DNA heteroduplex bound inside the central channel. The structural comparison of AsCpf1 with Cas9, a type II CRISPR-Cas nuclease, reveals both striking similarity and major differences, thereby explaining their distinct functionalities. AsCpf1 contains the RuvC domain and a putative novel nuclease domain, which are responsible for cleaving the non-target and target strands, respectively, and for jointly generating staggered DNA double-strand breaks. AsCpf1 recognizes the 5'-TTTN-3' protospacer adjacent motif by base and shape readout mechanisms. Our findings provide mechanistic insights into RNA-guided DNA cleavage by Cpf1 and establish a framework for rational engineering of the CRISPR-Cpf1 toolbox.
Organizational Affiliation:
Department of Biological Sciences, Graduate School of Science, The University of Tokyo, Tokyo 113-0032, Japan.