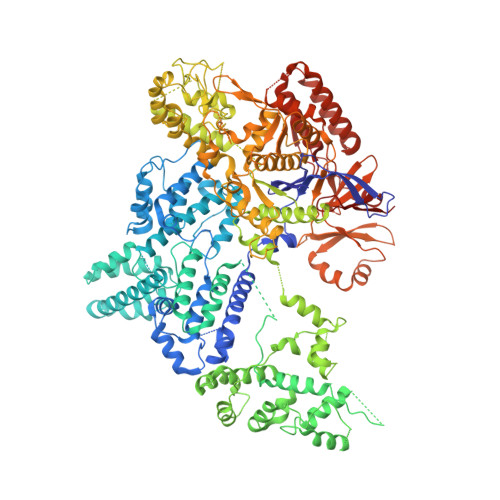
Structures of Cas9 Endonucleases Reveal RNA- Mediated Conformational Activation
Jinek, M., Jiang, F., Taylor, D.W., Sternberg, S.H., Kaya, E., Ma, E., Anders, C., Hauer, M., Zhou, K., Lin, S., Kaplan, M., Iavarone, A.T., Charpentier, E., Nogales, E., Doudna, J.A.(2014) Science 343: 47997
- PubMed: 24505130
- DOI: https://doi.org/10.1126/science.1247997
- Primary Citation of Related Structures:
4CMP, 4CMQ, 4OGC, 4OGE - PubMed Abstract:
Type II CRISPR (clustered regularly interspaced short palindromic repeats)-Cas (CRISPR-associated) systems use an RNA-guided DNA endonuclease, Cas9, to generate double-strand breaks in invasive DNA during an adaptive bacterial immune response. Cas9 has been harnessed as a powerful tool for genome editing and gene regulation in many eukaryotic organisms. We report 2.6 and 2.2 angstrom resolution crystal structures of two major Cas9 enzyme subtypes, revealing the structural core shared by all Cas9 family members. The architectures of Cas9 enzymes define nucleic acid binding clefts, and single-particle electron microscopy reconstructions show that the two structural lobes harboring these clefts undergo guide RNA-induced reorientation to form a central channel where DNA substrates are bound. The observation that extensive structural rearrangements occur before target DNA duplex binding implicates guide RNA loading as a key step in Cas9 activation.
Organizational Affiliation:
Department of Biochemistry, University of Zurich, CH-8057 Zurich, Switzerland.