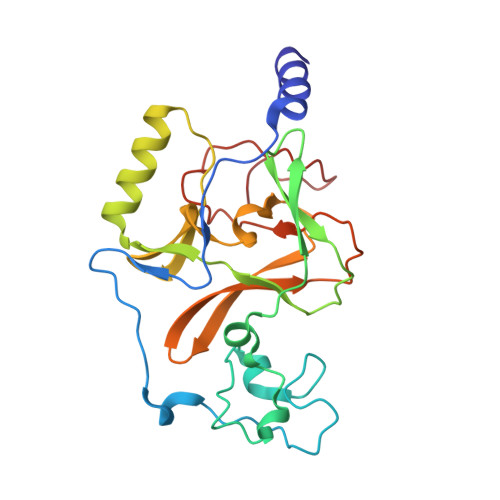
The structure of NSD1 reveals an autoregulatory mechanism underlying histone H3K36 methylation
Qiao, Q., Li, Y., Chen, Z., Wang, M., Reinberg, D., Xu, R.M.(2010) J Biol Chem 286: 8361-8368
- PubMed: 21196496
- DOI: https://doi.org/10.1074/jbc.M110.204115
- Primary Citation of Related Structures:
3OOI - PubMed Abstract:
The Sotos syndrome gene product, NSD1, is a SET domain histone methyltransferase that primarily dimethylates nucleosomal histone H3 lysine 36 (H3K36). To date, the intrinsic properties of NSD1 that determine its nucleosomal substrate selectivity and dimethyl H3K36 product specificity remain unknown. The 1.7 Å structure of the catalytic domain of NSD1 presented here shows that a regulatory loop adopts a conformation that prevents free access of H3K36 to the bound S-adenosyl-L-methionine. Molecular dynamics simulation and computational docking revealed that this normally inhibitory loop can adopt an active conformation, allowing H3K36 access to the active site, and that the nucleosome may stabilize the active conformation of the regulatory loop. Hence, our study reveals an autoregulatory mechanism of NSD1 and provides insight into the molecular mechanism of the nucleosomal substrate selectivity of this disease-related H3K36 methyltransferase.
Organizational Affiliation:
From the National Laboratory of Biomacromolecules, Institute of Biophysics, Chinese Academy of Sciences, 15 Datun Road, Chaoyang District, Beijing 100101, China and.