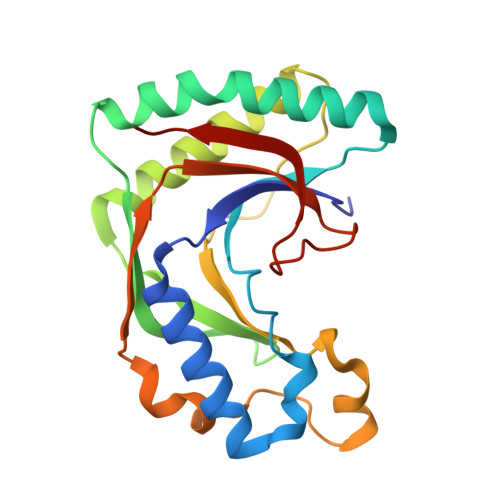
Akap18 Contains a Phosphoesterase Domain that Binds AMP
Gold, M.G., Smith, F.D., Scott, J.D., Barford, D.(2008) J Mol Biol 375: 1329
- PubMed: 18082768
- DOI: https://doi.org/10.1016/j.jmb.2007.11.037
- Primary Citation of Related Structures:
2VFK, 2VFL, 2VFY - PubMed Abstract:
Protein kinase A anchoring proteins (AKAPs), defined by their capacity to target the cAMP-dependent protein kinase to distinct subcellular locations, function as molecular scaffolds mediating the assembly of multicomponent complexes to integrate and organise multiple signalling events. Despite their central importance in regulating cellular processes, little is known regarding their diverse structures and molecular mechanisms. Here, using bioinformatics and X-ray crystallography, we define a central domain of AKAP18 delta (AKAP18(CD)) as a member of the 2H phosphoesterase family. The domain features two conserved His-x-Thr motifs positioned at the base of a groove located between two lobes related by pseudo 2-fold symmetry. Nucleotide co-crystallisation screening revealed that this groove binds specifically to adenosine 5'-monophosphate (5'AMP) and cytosine 5'-monophosphate (5'CMP), with the affinity constant for AMP in the physiological concentration range. This is the first example of an AKAP capable of binding a small molecule. Our data generate two functional hypotheses for the AKAP18 central domain. It may act as a phosphoesterase, although we did not identify a substrate, or as an AMP sensor with the potential to couple intracellular AMP levels to PKA signalling events.
Organizational Affiliation:
Section of Structural Biology, Institute of Cancer Research, Chester Beatty Laboratories, 237 Fulham Road, London SW3 6JB, UK.