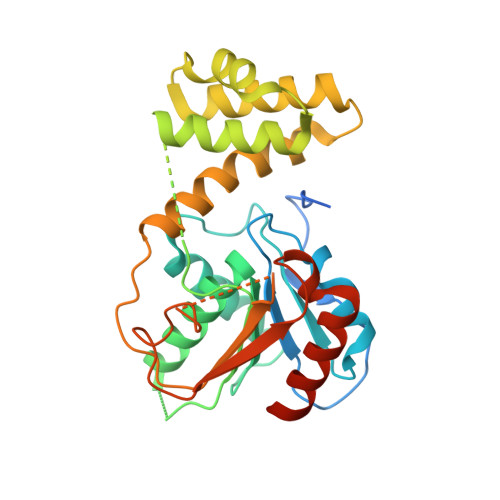
Crystal structure of the thioesterase domain of human fatty acid synthase inhibited by Orlistat.
Pemble, C.W., Johnson, L.C., Kridel, S.J., Lowther, W.T.(2007) Nat Struct Mol Biol 14: 704-709
- PubMed: 17618296
- DOI: https://doi.org/10.1038/nsmb1265
- Primary Citation of Related Structures:
2PX6 - PubMed Abstract:
Human fatty acid synthase (FAS) is uniquely expressed at high levels in many tumor types. Pharmacological inhibition of FAS therefore represents an important therapeutic opportunity. The drug Orlistat, which has been approved by the US Food and Drug Administration, inhibits FAS, induces tumor cell-specific apoptosis and inhibits the growth of prostate tumor xenografts. We determined the 2.3-A-resolution crystal structure of the thioesterase domain of FAS inhibited by Orlistat. Orlistat was captured in the active sites of two thioesterase molecules as a stable acyl-enzyme intermediate and as the hydrolyzed product. The details of these interactions reveal the molecular basis for inhibition and suggest a mechanism for acyl-chain length discrimination during the FAS catalytic cycle. Our findings provide a foundation for the development of new cancer drugs that target FAS.
Organizational Affiliation:
Center for Structural Biology and Department of Biochemistry, Wake Forest University School of Medicine, Medical Center Boulevard, Winston-Salem, North Carolina 27157, USA.