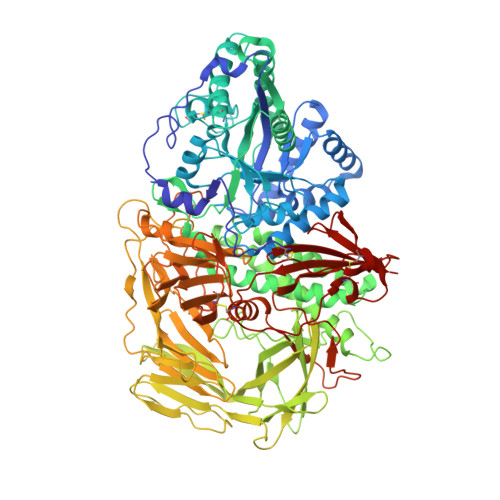
Structure of Golgi alpha-mannosidase II: a target for inhibition of growth and metastasis of cancer cells.
van den Elsen, J.M., Kuntz, D.A., Rose, D.R.(2001) EMBO J 20: 3008-3017
- PubMed: 11406577
- DOI: https://doi.org/10.1093/emboj/20.12.3008
- Primary Citation of Related Structures:
1HTY, 1HWW, 1HXK - PubMed Abstract:
Golgi alpha-mannosidase II, a key enzyme in N-glycan processing, is a target in the development of anti- cancer therapies. The crystal structure of Drosophila Golgi alpha-mannosidase II in the absence and presence of the anti-cancer agent swainsonine and the inhibitor deoxymannojirimycin reveals a novel protein fold with an active site zinc intricately involved both in the substrate specificity of the enzyme and directly in the catalytic mechanism. Identification of a putative GlcNAc binding pocket in the vicinity of the active site cavity provides a model for the binding of the GlcNAcMan(5)GlcNAc(2) substrate and the consecutive hydrolysis of the alpha1,6- and alpha1,3-linked mannose residues. The enzyme-inhibitor interactions observed provide insight into the catalytic mechanism, opening the door to the design of novel inhibitors of alpha-mannosidase II.
Organizational Affiliation:
Ontario Cancer Institute and Department of Medical Biophysics, University of Toronto, 610 University Avenue, Toronto, Ontario, Canada.